Research
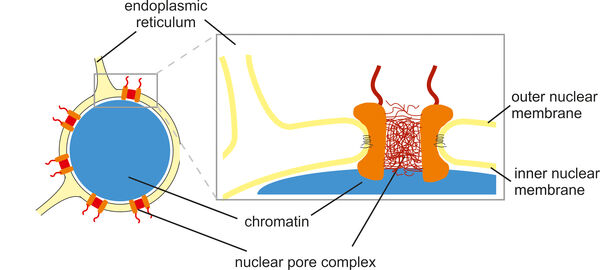
The cell nucleus is surrounded by a nuclear envelope that separates the cytoplasm from the interior of the nucleus. This separation requires efficient and selective transport gates that connect the sites of gene transcription and protein translation. Nuclear pore complexes fulfill this function by enabling the transport of proteins and nucleic acids while excluding inert molecules from passage. In addition, nuclear pore proteins are involved in gene regulation, which explains why many of them are implicated in the pathogenesis of malignant tumors as well as other human diseases such as atrial fibrillation, premature aging and muscle or lipodystrophies. In many cases, the underlying molecular disease mechanisms are not yet fully understood.
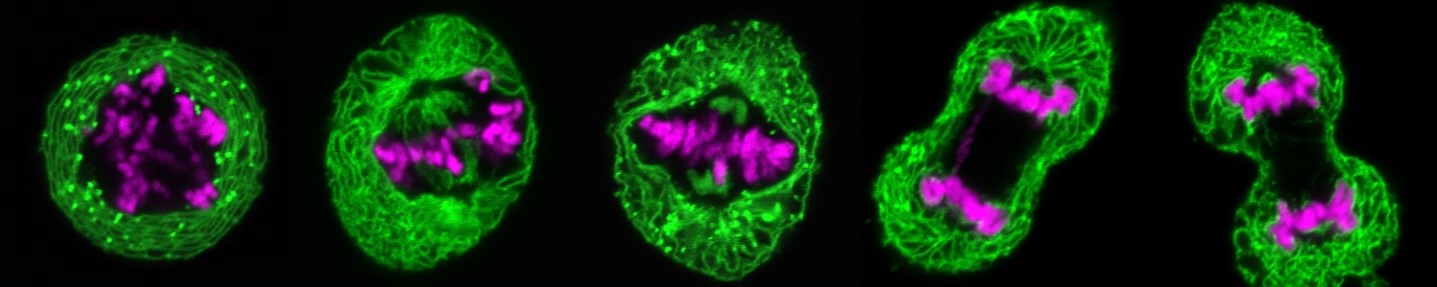
During cell division, the nuclei of human and animal cells undergo remarkable morphological and functional changes: At the beginning of mitosis, the nuclear pore complexes as well as the nuclear envelope disintegrate, while the chromatin condenses to allow the formation of the mitotic spindle, which distributes the DNA to the two nascent daughter cells. At the end of mitosis, the interphase structure of the nucleus must be restored to ensure the ability for DNA replication and transcription. This includes the decondensation of the highly condensed mitotic chromatin, the formation of a new closed nuclear envelope around the decondensing chromatin and the simultaneous reconstruction of the nuclear pore complexes and their integration into the newly forming nuclear envelope. All this happens in parallel and apparently well-coordinated within a short time window in the cell cycle. Many of the factors involved and their molecular mechanisms - particularly with regard to regulation and coordination - are at best only rudimentarily understood.
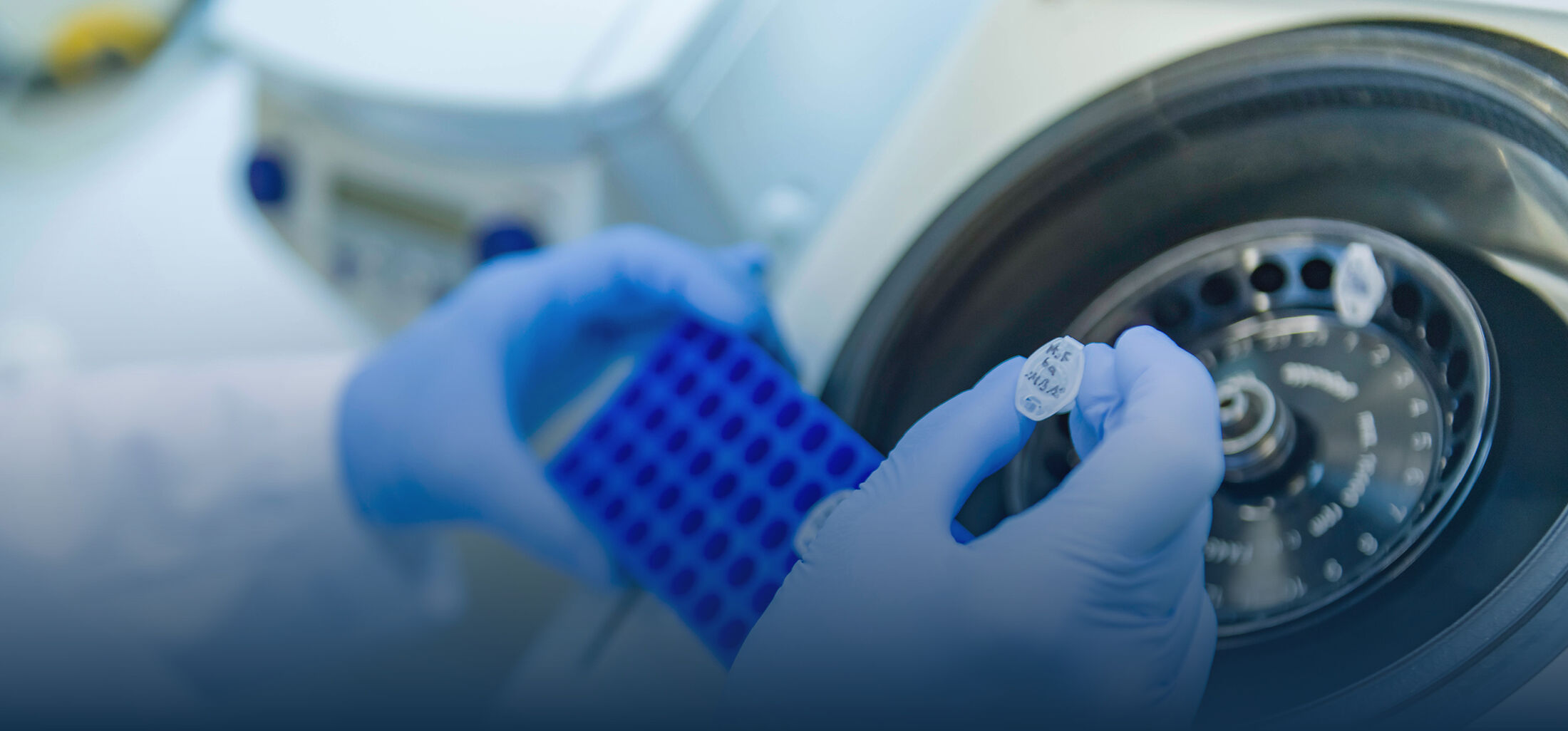
We investigate various aspects of nuclear remodeling in order to understand these processes in their cellular complexity, to decipher molecular mechanisms and to generate a detailed picture of the relevant factors and their interactions. To this end, we use a variety of biochemical (e.g. protein-protein interaction assays, membrane binding assays, enzyme kinetics) and cell biological experiments (e.g. live cell imaging, cell transfections as well as knock-outs and knock-downs). In particular, we close the methodological gap between well-defined minimalistic biochemical experiments and very complex investigations in living cells. In addition to mammalian cells, we also use freshly spawned frog eggs to obtain cell extracts for various cell-free reconstitution experiments. In these experiments, highly complex cellular processes, such as chromatin decondensation or the reconstruction of the nuclear envelope at the end of mitosis, can be tailor-made in the test tube and their molecular mechanisms analyzed.
Nuclear pore complexes
With a size of up to 125 MDa, nuclear pore complexes are among the largest protein complexes in cells. Their stepwise and coordinated assembly from over five hundred individual molecules and their integration into the nuclear envelope are impressive examples of molecular self-organization. We analyze this process using biochemical and cell biological methods, including cell-free assays and live cell imaging. We define the role of transmembrane proteins within nuclear pore complexes and investigate how nuclear pore proteins often interact with nuclear envelope membranes via amphipathic helices. This interaction is crucial for the strong curvature of the nuclear membranes into a pore structure.
While many nuclear pore proteins form a diffusion barrier in the center of the nuclear pore to prevent transport of most proteins, a few enable transport by establishing an activity gradient of the small GTPase Ran at the nuclear envelope. We are investigating how the nuclear pore protein Nup50 stimulates nuclear transport by activating Ran on the nuclear side of the pore, binding it and making it available for the transport process.
Some mutations in nucleoporins are associated with hereditary diseases. For example, we are investigating how such mutations can lead to nephrotic syndrome in children or cause amyotrophic lateral sclerosis. We are analyzing whether they cause defects in the assembly of nuclear pores or malfunctions in nuclear transport pathways and why these mutations have particularly serious effects in kidney or nerve cells.
Further reading:
Kutay U, Jühlen R, Antonin W. (2021). Mitotic disassembly and reassembly of nuclear pore complexes. Trends in Cell Biology S0962-8924(21)00139-2. doi: 10.1016/j.tcb.2021.06.011
Hamed M & Antonin W. (2021). Dunking into the Lipid Bilayer: How Direct Membrane Binding of Nucleoporins Can Contribute to Nuclear Pore Complex Structure and Assembly. Cells. 10(12):3601. doi: 10.3390/cells10123601.
Holzer G, De Magistris P, Gramminger C, Sachdev R, Magalska A, Schooley A, Scheufen A, Lennartz B, Tatarek-Nossol M, Lue H, Linder MI, Kutay U, Preisinger C, Moreno-Andrés D, Antonin W. (2021). The nucleoporin Nup50 activates the Ran guanine nucleotide exchange factor RCC1 to promote NPC assembly at the end of mitosis. EMBO Journal e108788. doi: 10.15252/embj.2021108788
Braun DA, Sadowski CE, Kohl S, Lovric S, Astrinidis SA, Pabst WL, Gee HY, Ashraf S, Lawson JA, Shril S, Airik M, Tan W, Schapiro D, Rao J, Choi WI, Hermle T, Kemper MJ, Pohl M, Ozaltin F, Konrad M, Bogdanovic R, Büscher R, Helmchen U, Serdaroglu E, Lifton RP, Antonin W, Hildebrandt F (2016). Mutations in nuclear pore genes NUP93, NUP205 and XPO5 cause steroid-resistant nephrotic syndrome. Nature Genetics, 48 (4): 457-465.
Nuclear envelope dynamics
At the beginning of mitosis, the nuclear envelope disintegrates so that its protein and lipid components are absorbed into the endoplasmic reticulum. At the end of mitosis, a new nuclear envelope forms from the endoplasmic reticulum in each developing daughter cell, enclosing the chromatin. We are investigating which proteins of the nuclear envelope and which changes to the chromatin are required to dissolve and rebuild the nuclear envelope. In particular, we use high-resolution live cell imaging in combination with quantitative, automated image analysis to capture dynamic cellular processes with temporal and spatial precision. To this end, we are developing open-source software tools based on deep learning. These tools enable us to automatically detect the different phases of mitosis and efficiently analyze large amounts of data generated during live cell imaging. This allows us to extract quantitative parameters and visualize them clearly. In this way, we are able to precisely identify defects during mitosis and in other phases of the cell cycle.
Further reading:
De Magistris P and Antonin W (2018). The Dynamic Nature of the Nuclear Envelope. Current Biology 28(8): R487–R497
Moreno-Andrés D, Bhattacharyya A, Scheufen A, Stegmaier J (2022). LiveCellMiner: A new tool to analyze mitotic progression. PLoS One 7;17(7):e0270923. doi: 10.1371/journal.pone.0270923
Jose A, Roy R, Moreno-Andrés D, Stegmaier J (2024). Automatic detection of cell-cycle stages using recurrent neural networks. PLoS One. 19(3):e0297356. doi: 0.1371/journal.pone.0297356.
Scheufen A & Moreno-Andrés D (2025). Quantitative Live-Cell Imaging to Study Chromatin Segregation and Nuclear Reformation. Methods Mol Biol . 2025:2874:47-60. doi: 10.1007/978-1-0716-4236-8_5
Chromatin dynamics in mitosis
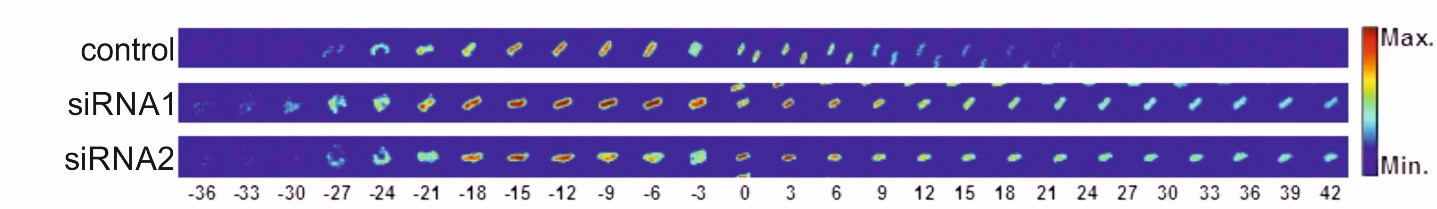
In human cells, mitotic chromatin is up to fifty times more condensed than in interphase. At the end of mitosis, it decondenses to restore the state required for transcription and replication. Although this process is crucial for cell division, the factors involved are largely unknown.
We are investigating which factors are involved in chromatin decondensation and which reactions they mediate. We also analyze which chromatin modifications, including histone and DNA modifications, are required during decondensation and how these processes are functionally and regulatory related to nuclear envelope formation and nuclear pore complexes. Since RNA-protein complexes and RNA helicases play an essential role in decondensation on the surface of mitotic chromosomes, we focus in particular on the analysis of these RNA-protein complexes and on which RNAs are primarily involved in this process. For our investigations, we use cell-free assays and experiments in mammalian cells, including RNAi and inhibitor-based screenings as well as live cell imaging.

Further reading:
Antonin W and Neumann H (2016). Chromosome condensation and decondensation during mitosis. Current Opinion in Cell Biology, 40:15-22.
Magalska, A, Schellhaus, AK, Moreno-Andres, D, Zanini, F, Schooley, A, Sachdev, R, Schwarz, H, Madlung, J, Gerken, J and Antonin, W (2014). RuvB-like ATPases function in chromatin decondensation at the end of mitosis. Developmental Cell, 31 (3): 305-318.
Jühlen R, Wiesmann S, Scheufen A, Stausberg T, Braun I, Strobel C, Llera-Brandt C, Rappold S, Suluyala R, Tatarek-Nossol M, Lennartz B, Lue H, Schneider M, Perez-Correa F, Moreno-Andres D, and Antonin W (2025). The DEAD-box helicase eIF4A1/2 acts as RNA chaperone during mitotic exit enabling chromatin decondensation. Nat Comm doi: 10.1038/s41467-025-57592-1.



